 
At time zero, the survival probability is 1.0 (or 100% of the participants are alive).The vertical tick mark on the curves means that a patient was censored at this time. A vertical drop in the curves indicates an event. The lines represent survival curves of the two groups. The horizontal axis (x-axis) represents time in days, and the vertical axis (y-axis) shows the probability of surviving or the proportion of people surviving. = "hv", # add the median survival pointer.Ĭ("Male", "Female"), # change legend labels.Ĭ("#E7B800", "#2E9FDF") # custom color palettes. ot = TRUE, # plot the number of censored subjects at time t text = FALSE,# show bars instead of names in text annotations l = T,# colour risk table text annotations. 
Risk.table = "abs_pct", # absolute number and percentage at risk. Ggtheme = theme_light(), # customize plot and risk table with a theme. Xlab = "Time in days", # customize X axis label.ī = 200, # break X axis in time intervals by 200. Pval = TRUE, # show p-value of log-rank test.Ĭonf.int = TRUE, # show confidence intervals forĬ = "step", # customize style of confidence intervals legend.labs to change the legend labels.įit, # survfit object with calculated statistics.As suggested by Marcin Kosinski, This is a good additional feedback to survival curves, so that one could realize: how do survival curves look like, what is the number of risk set AND what is the cause that the risk set become smaller: is it caused by events or by censored events? ot = TRUE to plot the number of censored subjects at time t..l = TRUE and .text = FALSE to provide bars instead of names in text annotations of the legend of risk table.risk.table = “abs_pct”to show both absolute number and percentage of individuals at risk.= 200 break x axis in time intervals by 200. 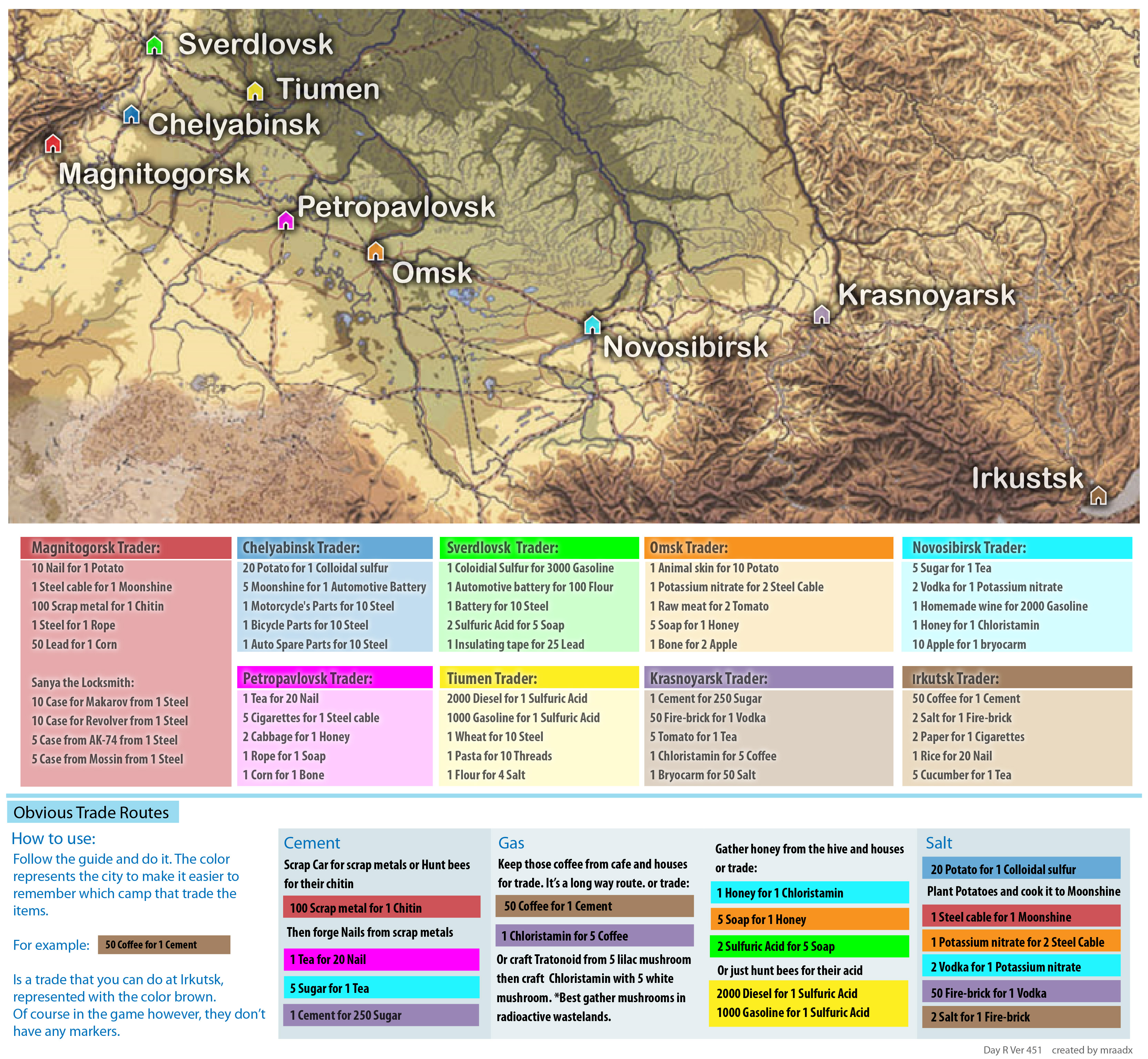
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
